
2. Spatial pre-processing
2.1 Realignment (spm_realign_ui.m)
Realignment of an image time-series of the same modality.

Specify:
![]()
Specify (for fMRI only):
![]()
Select scans:
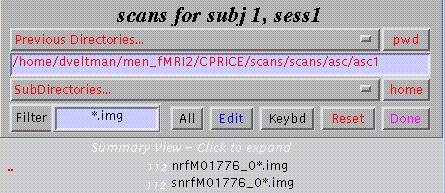
Clicking the *.img will select those images, clicking the preceding number (in white) will show individual scans. SPM displays the total number of selected scans (lower part of window).
Other options:
Filter disables filter
All selects all images shown
Edit allows editing of image file names
Keybd allows command line input of image file names
Reset will undo selection
Done completes selection.
Specify:

Coregister only: determines parameters used to realign to the first image in the selected series. For multiple sessions, the first scan of each session is realigned to the first scan of the first session; then the images within each session are realigned to the first scan of the session. Saves the transform parameters for each filename.img in filename.mat. Realignment parameters are saved for each session as realignment_params_*.txt, which can be used as a set of 6 covariates to regress out movement-related activations.
N.B.
The first image = the first selected image. This need not be the first acquired in time.
For each realignment, the current *.mat files are used and updated.
All output images are written to the same directory as the input images. The mean*.img is written to the directory containing the images for the first session.
In PET, realignment is in two steps: all images are realigned to the first and a mean image is computed to which all images are realigned in a second pass.
Reslice only: reslices and saves images using the transforms specified by *.mat files. Transforms filename.img using filename.mat and saves as rfilename.img.
*Coregister & Reslice (*=default): above steps combined. Saves transform parameters for filename.img in filename.mat. Reslices and saves images as rfilename.img.
Specify:
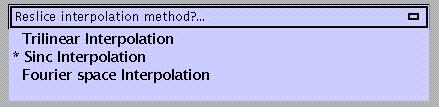
Trilinear interpolation: uses trilinear interpolation to resample images.
Sinc interpolation (default): uses a truncated (kernel is 9x9x9 voxels) sinc interpolation. Slower than trilinear but more accurate; recommended for fMRI time-series.
Fourier space interpolation: added option also found in other packages (AIR). Performs rigid body rotations as shears in Fourier space (Eddy et al. 1996, MRM 36, 923-931). For isotropic voxels only.
Specify:
All images (1-n): reslices all images including the first which is untransformed (i.e., SPM duplicates the first and saves it as rfirstimage).
Images 2-n: reslices all images except the first (useful when the first image has a different modality).
All images + Mean Image (default): reslices all images and creates a mean of all resliced images, which is saved as meanfilename.img.
Mean Image only: creates mean image only.
N.B.
Because each reslicing slightly degrades the images, this may be postponed until the normalisation step. It is useful, however, to create a mean image to determine parameters from at the normalisation stage (2.4) or to coregister to a structural MR (2.3). Select Coregister & Reslice, Trilinear or Sinc interpolation (trilinear is probably sufficient and much faster) and Mean Image Only. The other images will have only their *.mat files updated.
If the ORIGIN coordinates in the image headers need adjustment, this should be done prior to realignment (because SPM will ignore the ORIGIN coordinates in the header once a *.mat file is created). Select the Display option (lower panel), select an image, and place the crosshair at the anterior commissure. Select HDRedit, select 'set Origin', and copy & paste the coordinates (in voxels) from the Graphics window. Apply to all time-series images (see 5 for discussion of utilities).
Specify (for fMRI only):
![]()
This removes interpolation errors arising from reslicing of the data by doing a regularised fit for motion-related variance and removes estimated variance from resliced images. However, if there is any motion-related variance that is correlated with the task, this option may remove variance of interest (i.e. true positives).
Output from spm_realign:
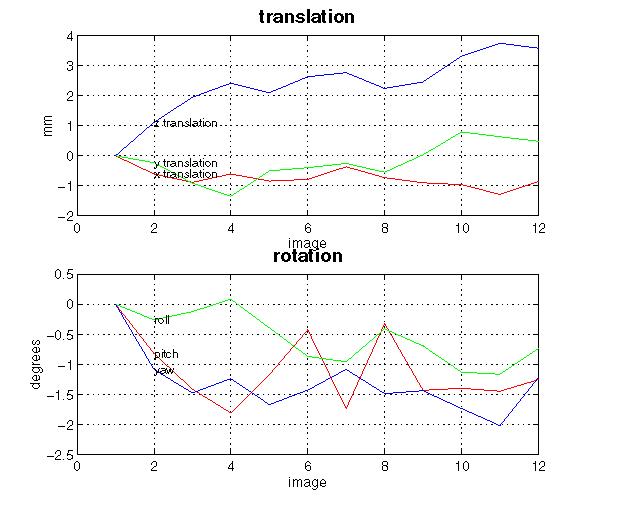
Display (shown here for a 12-scan PET study): spm_realign displays realignment parameters in graphic form. This display is written to the spm99.ps file in the working directory. This file can be viewed using a postscript reader package such as ghostview or pageview.
Files: the realigned images r*.img, r*.hdr, and the updated .mat files are written to the input file directory. The mean*.img and mean*.hdr are written to the input file directory for the first session. Realignment parameters are saved as realignment_params_*.txt to the input file directory for each session.
Customisations for realignment (optional):
N.B.
Changes in default settings are valid only during one session. After quitting and restarting SPM, these adjustments need to be renewed.
From the 'Defaults' options (lower panel), select 'Realignment':
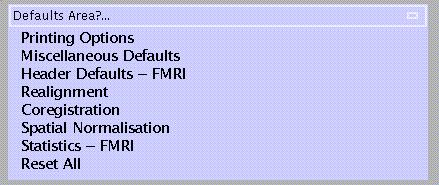
Specify:

allow separate coregistration and reslicing (default)
always combine coregistration and reslicing
Specify:
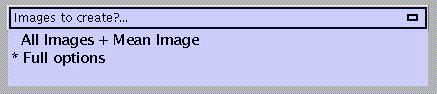
always create all images & mean image
full options (default, see above)
Specify:
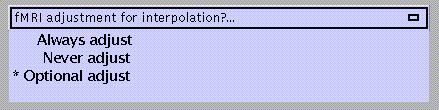
always adjust for interpolation errors
never adjust for interpolation errors (e.g., when realignment parameters are used as covariates)
optional adjust (default)
Specify:

always mask images (=default): to avoid movement-related artefactual variance at the edge of the FOV
optional mask
Specify:
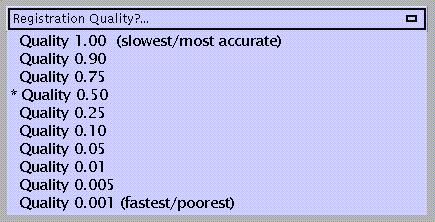
which refers to the density of voxel sampling for computation of realignment parameters. The quality improves with higher density, which is also more time-consuming.
2.2 Coregistration (spm_coreg_ui.m)
Between and within modality image coregistration.

Specify:
![]()
Specify:

Coregister only: determines parameters used to coregister to one image to another. This option saves the transform parameters filename.img in filename.mat.
Reslice only: reslices and saves images using the transforms specified by the *.mat files. Transforms filename.img using filename.mat and saves as rfilename.img.
*Coregister & Reslice (*=default): above steps combined. Saves transform parameters for filename.img in filename.mat. Reslices and saves images as rfilename.img.
FOR 'Coregister only' AND 'Coregister & Reslice':
Specify:
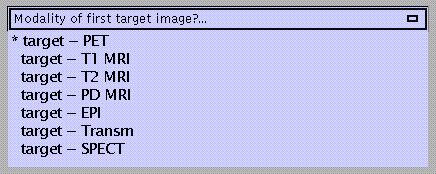
Specify:
Select:
![]()
Select:
![]()
Select (or press 'Done'):
![]()
The TARGET image is the image to which the OBJECT image is realigned. OTHER images (if any) are transformed in the same way as the OBJECT image.
For example, to realign a structural T1-MRI to a fMRI time-series:
TARGET : mean EPI
OBJECT : T1
OTHER : -
To realign a sequence of PET-images to a structural T1-MRI:
TARGET: T1
OBJECT: mean PET
OTHER: PET1.img, PET2.img,?.
N.B.
If the target and object images have different modalities, an image segmentation (see 2.6) is performed to partition target and object into CSF, white and grey matter. These partitions are then used in the coregistration.
FOR 'Reslice Only':
Select:
![]()
The space of this image (i.e. the space defined by the .mat file associated with this image) will be used to reslice the object image.
Select:
![]()
This image is the object image that will be resliced in the space of the previously selected image. The image will be resliced at the voxel size and origin of the previously selected image and will have the same space defined by the corresponding .mat file.
Output from spm_coregister:
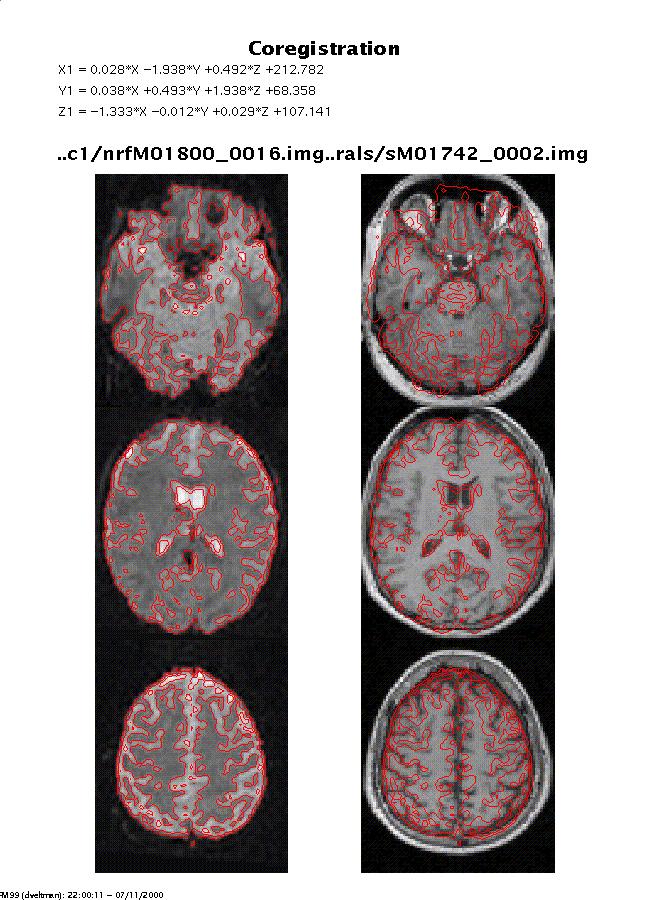
Display: spm_coregister displays the coregistered images with the contours of the OBJECT image overlaid on both sets of images. This display is written to the spm99.ps file in the working directory. If OBJECT and TARGET image have different modalities, a segmentation is also carried out for both images. This is displayed as in spm_segment (see 2.6) and written to the spm99.ps file.
Files: the coregistered images rfilename.img and rfilename.hdr are written to the same directory as the input images if 'reslice' is specified. The current rfilename.mat files are updated.
Customisations for coregistration (optional):
N.B.
Changes in default settings are valid only during one session. After quitting and restarting SPM, these adjustments need to be renewed.
From the 'Defaults' options (lower panel), select 'coregistration':
Specify:

use default registration
use mutual information registration (advisable when default between-modality coregistration doesn't work due to e.g. poor grey/white contrast, so that the segmentation step fails)
Specify (for default between-modality coregistration only):

do all three coregistration steps (i.e., (1) 6-parameter (rigid body) normalisation of object and target image to a template of its own modality, (2) segmentation of object and target image, (3) coregistering the segmented images using the results of (1) as a starting point; =default)
do first coreg step only (6-parameter normalisation of object and target image, each to a template of its own modality), e.g. when coregistering two images with poor grey/white contrast such as a PET transmission scan and a CT-scan, or as a first step prior to mutual information registration
2.3 Slice timing (spm_slice_timing.m)
Correction for differences in slice acquisition time.

This is only for fMRI. Corrects for differences in acquisition time between slices during sequential imaging (as in echo-planar imaging). It is especially relevant for event-related designs. This routine performs a phase shift, resulting in each time series having the values that would have been obtained had the slice been acquired at (for example) the beginning of each TR.
N.B.
This step can be performed either before or after realignment (but before normalisation). Both procedures have potential disadvantages:
slice time correction before realignment may interpolate signals from different brain regions if there is significant head movement (which may be particularly problematic near the edge of the brain, where large signal differences occur across nearby voxels);
slice time correction after realignment however may shift voxels to adjacent slices (and hence different time points).
The latter problem is especially relevant for interleaved acquisitions, where the time difference between adjacent slices may be ½ TR. It is therefore recommended to perform slice time correction first for interleaved sequences.
The latter problem is less relevant to sequential (ascending/descending) acquisitions, where the time difference between adjacent slices is small, and so realignment may be better first for ascending/descending sequences, to allow for movement effects.
Specify:
![]()
Select:
![]()
Specify:

ascending (first slice = bottom; e.g. 1 2 3 ???..68)
descending (first slice = top; e.g. 68 67 66 ??..1)
interleaved (first slice = top; e.g. 68 66 64 ??2 67 65 63?.1)
user specified (e.g., other interleaved sequences; SPM will prompt for a vector specifying slice acquisition order, where the slices are numbered in Analyze format)
Specify:
![]()
SPM will suggest the middle slice (in space) as the default (in the example above: slice 34). This is particularly useful for ascending/descending sequences because the phase shift interpolation tends to become less accurate with increasing shifts, and slices further away from the middle slice tend to be of less interest. When the regions of interest do lie in extreme (top or bottom) slices however, the reference slice may be chosen to correspond to these regions, so that interpolation artefacts are minimised (the signal in the reference slice itself is unchanged by the interpolation).
N.B.
For fMRI analyses, SPM divides each TR into a number of time bins (by default this number=16). SPM by default then uses the first time bin as the reference or sampled bin. This bin provides the values from which the covariates are constructed.
When the first slice in time is NOT used as the reference slice during slice-timing correction, the default sampled bin (=1) must be adjusted prior to statistical analysis.
For example, if the middle slice was chosen as the reference slice for slice timing correction, the middle time bin must be set to the sampled time bin. If the number of time bins =16 for example, the sampled time bin should =8.
To change these settings, select Defaults, select 'Statistics - fMRI', set 'Sampled bin' to 8. This change must be made before creating the design matrix.
Specify:
![]()
Specify:
![]()
TR is defined as the time between the first slice of one scan and the first slice of the next scan; TA is the time between the first and the last slice within one scan. Without an interscan time delay, TA=TR-(TR/N), where N= number of slices (= default). When there is an interscan delay, TA must be adjusted.
N.B.
In this example, TR>4sec. Although slice timing correction is potentially more important for long TRs, interpolation also becomes less accurate. It is therefore advised to omit slice time correction for TR>3sec and model timing differences by including temporal derivatives as an additional regressor, or using a phase-invariant Fourier set (see 3.1).
Output from spm_slice_timing:
Files: afilename.hdr, afilename.img and afilename.mat files are written to the input file directory.
2.4 Normalisation (spm_sn3d.m)
Three-dimensional spatial normalisation of PET, SPECT and (f)MRI images to a standard space.

Normalisation consists of two steps: first the determination of an optimum 12-parameter (translations, rotations, zooms & shears) affine transformation (from an image to a template), followed by a nonlinear estimation of deformations. These parameters can be used to reslice other images coregistered to the image from which the parameters were determined.
Select:
Determine parameters only: determines parameters (linear and nonlinear) used to normalise image to a template. Saves normalisation parameters in filename_sn3d.mat.
Write normalised only: reslices and saves images using normalisation parameters specified by the selected sn3d.mat file. Applies parameters to filename.img (*.hdr, *.mat) and saves as nfilename.img (n*.hdr, n*.mat).
Determine parameters and write normalised (=default): the above steps combined. Saves normalisation parameters for filename.img in filename_sn3d.mat. Saves normalised images as nfilename.img (n*.hdr, n*.mat).
N.B.
All output images, headers and *.mat files as well as the *_sn3d.mat file are written to the input directory.
To assure optimal normalisation, image and template should be in a similar starting position. This can be tested with the Check Reg option in the lower panel. Use the Display option to reposition the image (see section 5).
Specify:
![]()
Select:
![]()
for example, a coregistered T1 structural image or a mean functional (PET or EPI) image.
Select:
![]()
Select:
![]()
The selected template image which should be the same modality (and have very similar contrast) as the image from which the parameters are to be determined.
N.B.
The space of the template images in SPM (referred to as MNI space) is based upon the Talairach system, but does not make assumptions about brain symmetry, and includes the cerebellum.
The stereotactic space is based on 152 brains from the Montréal Neurological Institute, and will eventually be replaced in due course by a 450-brain version for the entire ICBM consortium (http://www.loni.ucla.edu/ICBM ).
The 152 subject average brain was chosen rather than the MNI305 official standard brain because from the 152 subjects T2- and PD-weighted images were also available, allowing much more flexibility in the range of different MR contrasts that can be spatially normalized to the same stereotaxic space. See spm_templates.man for details.
A combination of templates may be used when the image to determine parameters from is different (e.g., intensity with a new MRI sequence) from the templates provided by SPM. For other purposes, e.g. normalising PET-ligand scans, creating a user-specified template should be considered.
Specify:

Nearest neighbour: fast but not very accurate and therefore not usually recommended.
Bilinear (=default): recommended for fMRI images which have been resliced at the realignment stage, and PET.
Sinc interpolation: uses a modified sinc interpolation (9x9x9 voxels kernel); slowest, recommended for fMRI data which have not been resliced at the realignment stage.
Output from spm_sn3d:
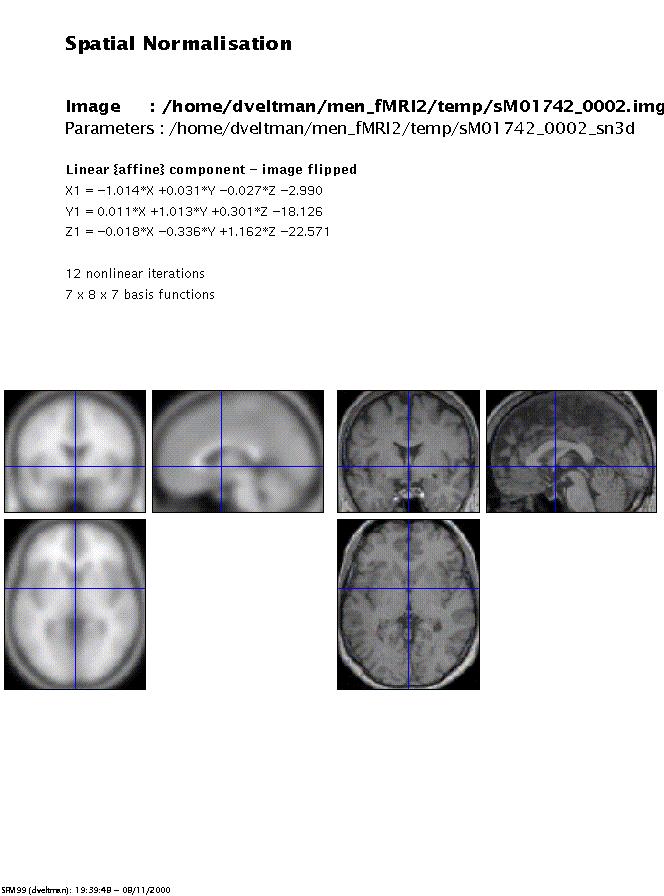
Display: spm_sn3d displays the normalised image and the template. The display is written to the spm99.ps file in the working directory.
Files: the normalisation parameters filename_sn3d.mat are written to the same directory as the input image(s). The normalised images nfilename.img (n*.hdr, n*.mat) are written to the input directory if 'write normalised' is specified.
Customisations for normalisation (optional):
N.B.
Changes in default settings are valid only during one session. After quitting and restarting SPM, these adjustments need to be renewed.
From the 'Defaults' options (lower panel), select 'Spatial Normalisation':
Specify:

Defaults for Parameter Estimation (first step)
Defaults for Writing Normalised (second step)
For changing Parameter Estimation defaults, specify:
Neurological Convention (R=R): SPM assumes that the images are in neurological convention (as used by SPM) so the starting estimates won't include a L-R flip (0 0 0 0 0 0 1 1 1 0 0 0)
Radiological Convention (L=R; default): SPM assumes that the images are in radiological convention so the starting estimates will include a L-R flip (0 0 0 0 0 0 -1 1 1 0 0 0)
Custom Starting Estimates:
![]()
enter these in the following order:
x translation (mm)
y translation (mm)
z translation (mm)
x rotation 'pitch' (radians)
y rotation 'roll' (radians)
z rotation 'yaw' (radians)
x scaling
y scaling
z scaling
x shear
y shear
z shear
N.B.
As an alternative to customising starting estimates, prior to normalisation the images may be repositioned using the Display option (see above).
Specify:
Allow customised (options for # nonlinear basis functions and iterations, type of regularisation, and masking)
Disallow customised normalisation (=default; standard options for # nonlinear basis functions and iterations, type of regularisation, and masking)
If customised is selected, specify:
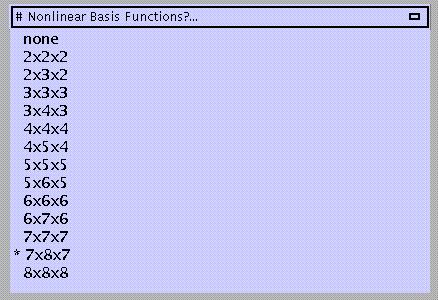
The default is 7x8x7 functions. A large number of functions will improve the quality of the normalisation provided the images do not contain gross lesions and/or have different contrasts compared to the template. More functions will be computationally slower, however.
Specify:
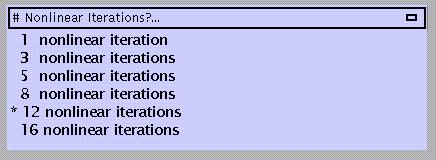
The default is 12 iterations. Again, the more iterations the better the result, but memory demands will similarly increase.
Specify:
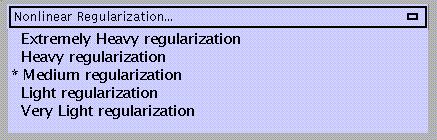
Regularisation involves stabilising the estimation of the deformations by penalising large deformations. Heavy regularisation basically means that more weight is applied to the penalty term. Therefore, if the spatial normalisation introduces excessive warping that is clearly wrong, more regularisation is needed. Conversely, if the images do not get warped enough to match the template, the amount of regularisation needs to be decreased. The default is medium regularisation.
Specify:
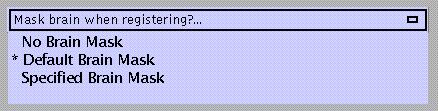
Masks template brain (1 1 1 1 .. for voxels within the brain, 0 0 0 0 .. for voxels outside the brain) in order to perform normalisation based on the shape of the brain instead of the skull. Not necessary when normalising a mean EPI image to the EPI template, in which case unmasked normalisation may give better results.
Specify:
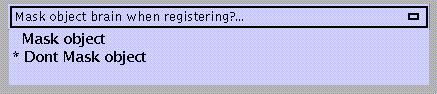
Similarly using 1 1 1 1 .. for voxels within the brain, 0 0 0 0 .. for voxels outside the brain. An object brain mask might be used e.g. to prevent an abnormal region (lesion) influencing the normalisation.
For changing Writing Normalised defaults, specify:
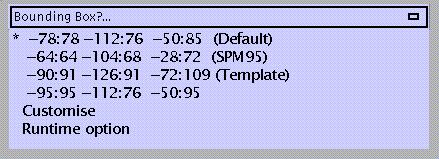
In mm distances (x y z) from the anterior commissure. When choosing 'Runtime option', SPM will prompt for bounding box dimensions during specification of the normalisation.
Specify:

In mm (x y z):
1 1 1
1.5 1.5 1.5
2 2 2 (=default; may lead to oversampling of the data; the advantage is that the data will require less smoothing to meet the assumptions of Gaussian field theory, disadvantage is that resulting *.img files will use more disk space)
3 3 3
4 4 4
1 1 2 (e.g., for structural MRI's with 1mm interslice gap)
2 2 4 (e.g., for EPI images with 2mm in-plane resolution and 2mm interslice gap)
Customise
Runtime option (i.e., SPM will prompt for voxel sizes during specification of the normalisation)
2.5 Smooth (spm_smooth.m)
Three-dimensional convolution of an image with a Gaussian kernel.

Specify:
![]()
Gaussian filter width. This can be a scalar (for isotropic voxels) or a vector of length 3 (sx sy sz) for anisotropic voxels. Suggested is 2-3 times the voxel size.
Select:
![]()
N.B.
Spatial smoothing has three important objectives:
it increases signal to noise ratio
it conditions the data so that they conform more closely to a Gaussian field model. This is important if Gaussian field theory is used to make statistical inferences about regionally specific effects (i.e., assign p-values)
it ensures that effects between different subjects are assessed on a reasonable spatial scale with respect to functional anatomy.
See also Matthew Brett's basic tutorial on smoothing at:
http://www.mrc-cbu.cam.ac.uk/Imaging/smoothing.html
Output from spm_smooth:
Images are saved as sfilename.img (s*.hdr, s*.mat) in the same directory as the input files.
2.6 Segment (spm_segment.m)
Segments MRI-image(s) into grey and white matter, CSF, and other.

Specify:
![]()
Select:
![]()
if more than one, they should be coregistered.
Specify:
![]()
If not, specify:
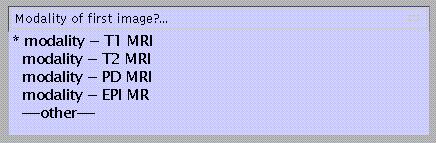
to match with a template which will be used for affine normalisation of the image(s). This is required for probabilistic segmentation.
Specify:
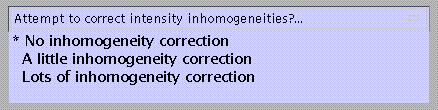

this option attempts to model intensity inhomogeneities using smooth basis functions.
If inhomogeneity correction is to be applied, specify:
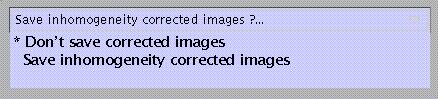
Inhomogeneity-corrected images may be saved as corr_filename.img for inspection.
Output from spm_segment.m:
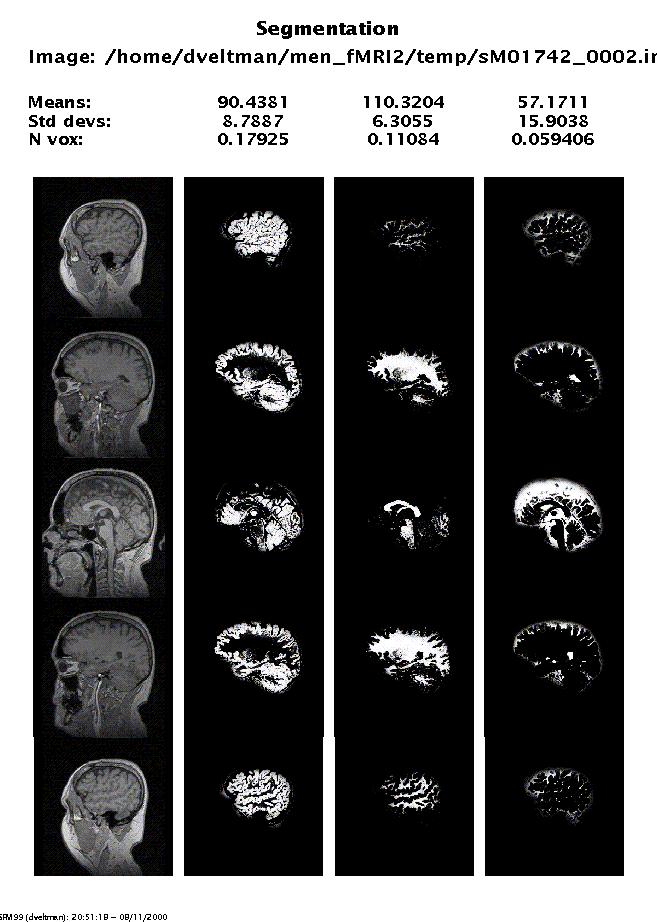
Display: spm_segment displays the segmentation results. This display is written to the spm.ps file in the working directory. N vox is the proportion of voxels belonging to each class, worked out from a subregion of the volume that encloses the brain.
Files: the segmented images are written as filename_seg1.img, filename_seg2.img, and filename_seg3.img for grey and white matter, and CSF. These images are saved to the input directory.